Research
We develop new methods and algorithms for electronic structure theory. Click the links below to find out more!
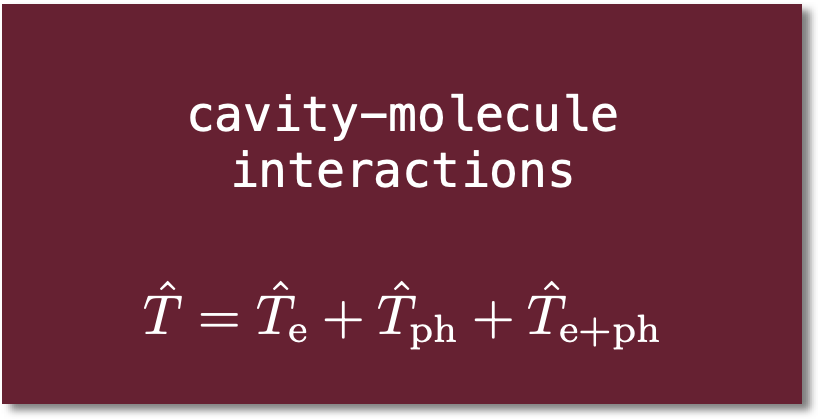



Local correlation: clusters in molecules (CIM)
By partitioning a large molecule into strongly-interacting domains, fragment-based local correlation approximations like the cluster in molecule (CIM) method overcome the steep polynomial scaling of CCSD(T) and other many-body methods. As system size increases, fragment sizes remain constant; the total cost of a computation increases only linearly with the number of fragments:
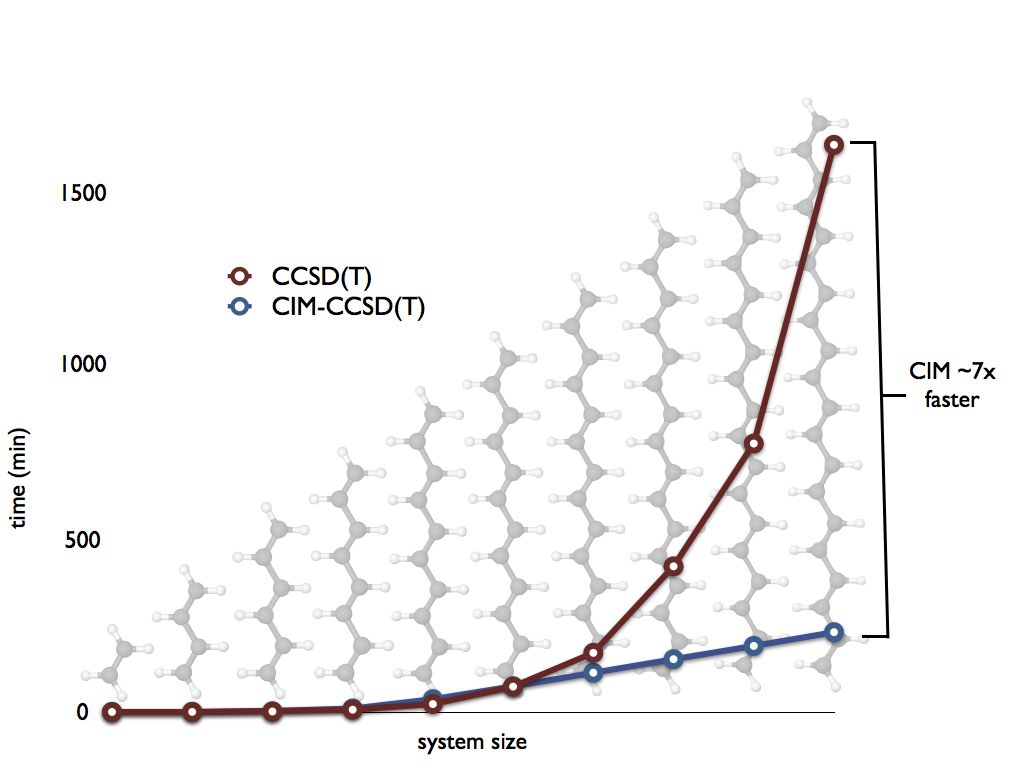
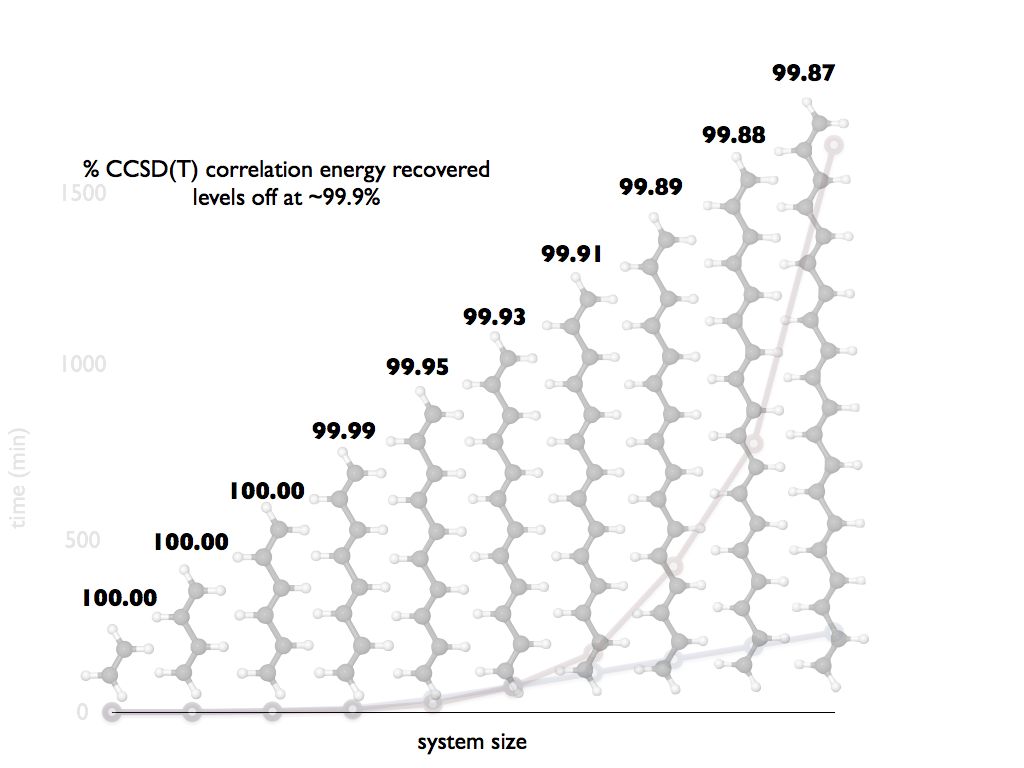
For many systems, we observe huge savings in time-to-solution with very little loss in accuracy. CIM methods often recover nearly 100% of the canonical correlation energy: