Our long-term goal is to reveal how variations in sequence and post-translational modifications of proteins modulate their structures and interactions. We are particularly interested in the biochemistry of the so-called “glycan shield” formed by viral spike glycoprotein complexes. Another area of long-term interest is the self-assembly of chemokines and how these assemblies are modulated through interactions with carbohydrates.
In pursuing these goals, we develop tandem-trapped ion mobility spectrometry/mass spectrometry instrumentation and novel computational approaches to elucidate protein structures from ion mobility data, and apply these methods to study human diseases.
Instrumentation: Development of tandem-trapped ion mobility spectrometry/mass spectrometry
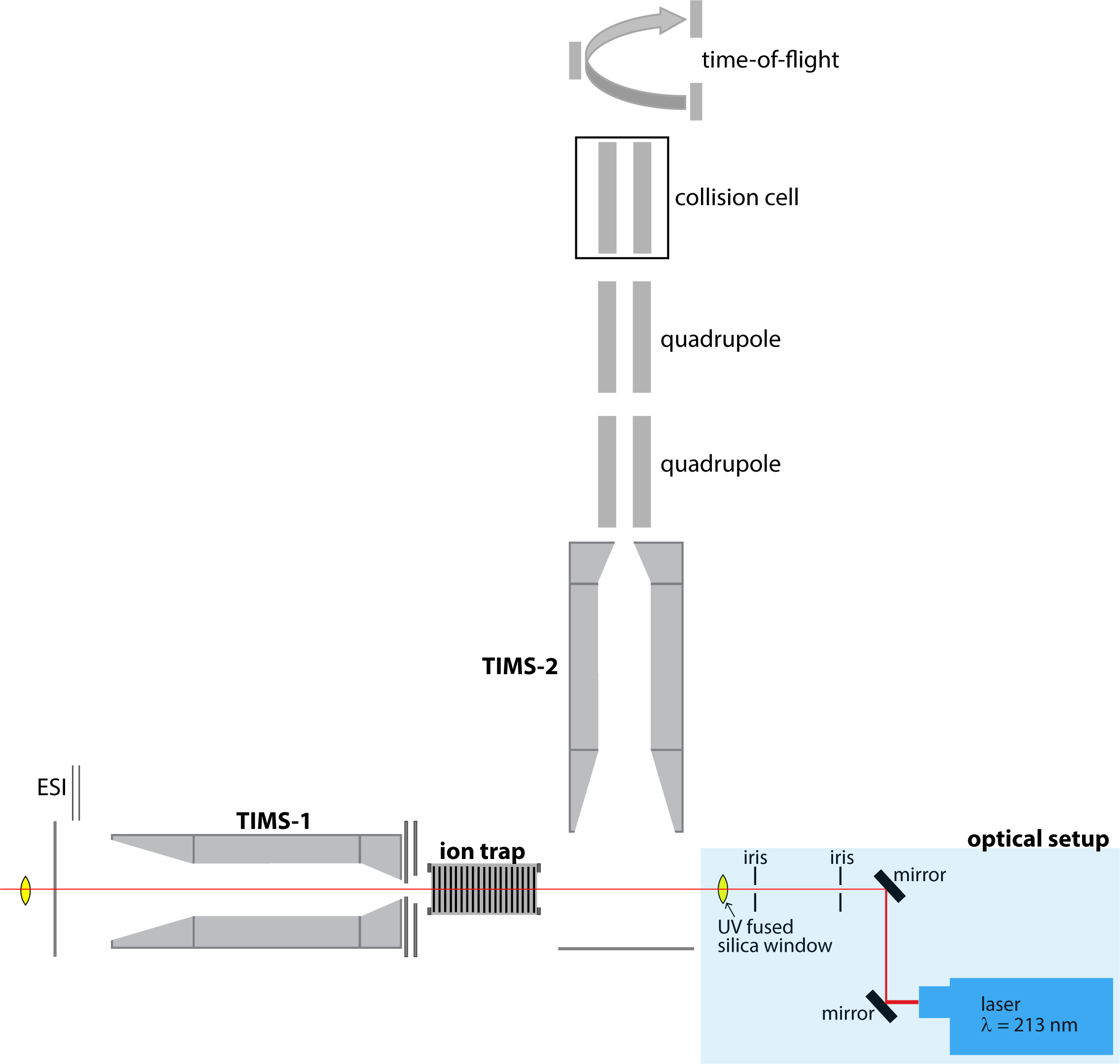
Our laboratory began working on trapped ion mobility spectrometry (TIMS) in 2014. Initially, we focused on the analytical characterization of the method itself as it pertains to biological applications. For example, we demonstrated that TIMS is suited for native mass spectrometry of protein systems (Analyst, 2016). Additionally, we developed an accurate, transferable calibration method for ion mobility measurements conducted in electrodynamic fields and non-stationary buffer gases (Analytical Chemistry 2018), which built on the prior derivation of a transport equation for TIMS from the kinetic theory of gases (International Journal of Mass Spectrometry, 2016). Subsequently, we pioneered the development of tandem-TIMS (Analyst 2018, US patent 2020) ad identified how tandem-TIMS can be used to study weakly-bound peptide assemblies (Journal of The American Society for Mass Spectrometry 2019 & Analytical Chemistry 2020). This development was followed by coupling of tandem-TIMS with UV photodissociation (Rapid Communications in Mass Spectrometry 2021) for top-down analysis of protein complexes. We further demonstrated proof-of-principle that tandem-TIMS is capable to characterize the primary, tertiary, and quaternary structures of glycoprotein complexes (Analytical Chemistry 2020).
In ongoing work, we apply and improve tandem-TIMS methods to characterize carbohydrate structures, and to identify how carbohydrates interact with proteins and how they modulate protein structure and function.
Selected publications:
- Liu, C., Ridgeway, M. E., Winfred, J. S. R. V., Polfer, N. C., Lee, J., Theisen, A., Wootton, C. A., Park, M. A. & Bleiholder, C. (2021). Tandem-trapped ion mobility spectrometry/mass spectrometry coupled with ultraviolet photodissociation. Rapid Communications in Mass Spectrometry, 35(22), e9192.
- Bleiholder, C., Liu, F. C., Chai, M. (2020). Comment on Effective Temperature and Structural Rearrangement in Trapped Ion Mobility Spectrometry. Analytical Chemistry, 92, 16329−16333.
- Park, M. A., Ridgeway, M. E., Bleiholder, C., & Liu, F. C. (2020). Tandem Ion Mobility Spectrometer. US Patent 10,794,861, Bruker Daltonics.
- Liu, F. C., Cropley, T. C., Ridgeway, M. E., Park, M. A., & Bleiholder, C. (2020). Structural Analysis of the Glycoprotein Complex Avidin by Tandem-Trapped Ion Mobility Spectrometry–Mass Spectrometry (Tandem-TIMS/MS). Analytical Chemistry, 92(6), 4459-4467.
- Liu, F. C., Ridgeway, M. E., Park, M. A., & Bleiholder, C. (2018). Tandem trapped ion mobility spectrometry. Analyst 143, 2249-2258.
- Chai, M., Young, M. N., Liu, F. C., & Bleiholder, C. (2018). A transferable, sample-independent calibration procedure for trapped ion mobility spectrometry (TIMS). Analytical Chemistry 90, 9040-9047.
- Kirk, S. R., Liu, F. C., Cropley, T., Carlock, H., & Bleiholder, C. (2019). On the preservation of non-covalent peptide assemblies in a tandem-trapped ion mobility spectrometer-mass spectrometer (TIMS-TIMS-MS). Journal of The American Society for Mass Spectrometry, 30, 1204-1212.
- Bleiholder, C. (2016). Towards measuring ion mobilities in non-stationary gases and non-uniform and dynamic electric fields (I). Equation of motion. International Journal of Mass Spectrometry 399-400, 1-9.
- Liu, F. C., Kirk, S. R., & Bleiholder, C. (2016). On the Structural Denaturation of Biological Analytes in Trapped Ion Mobility Spectrometry – Mass Spectrometry. Analyst 141, 3722-3730.
Theory: Structure-elucidation from ion mobility spectrometry/mass spectrometry
A major current challenge of applying ion mobility spectrometry to biological problems is to interpret the experimental data in terms of the molecular structure of the analyte. These challenges are rooted in the fact that ion mobility spectrometry provides no direct, atomic information about the investigated molecule. Instead, the structural information obtained from IM-MS is a mean “effective area” called momentum transfer cross section.
Our research initially focused on developing methods that enable rapid yet accurate cross section calculations of proteins and protein assemblies. To this end, we developed the Projection Superposition Approximation (PSA) method (International Journal of Mass Spectrometry, 2011) and the local collision probability approximation (LCPA, Analyst 2015) which significantly improves upon the PSA.
These methods make it fairly routine to accurately calculate cross sections for protein system in excess of ~100 kDa, providing the basis for biological applications of ion mobility spectrometry. However, algorithms that calculate a cross section for a given model structure do not by themselves decipher the protein structure from the experimental data. Hence, we developed the Structure Relaxation Approximation (SRA) for the elucidation of protein structures from ion mobility measurement (Journal of Physical Chemistry 2019). This is the first general method capable of predicting ion mobility spectra in order to interpret the experimentally measured spectra in terms of the protein structure. The underlying rationale for development of the SRA is that it allows predicting the “global” trends observed in the ion mobility experiments for various experimental conditions and charge states, thereby enabling detailed elucidation of the protein structure. For example, the SRA predicts that even the small protein ubiquitin largely retains its native inter-residue contacts with an intact hydrophobic core when studied by “soft” ion mobility measurements.
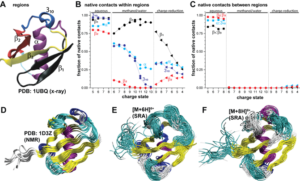
Selected publications:
- Bleiholder, C., Liu, F. C. & Chai, M. (2021). Calculation of momentum transfer cross-sections. Ion Mobility – Mass Spectrometry: Fundamentals and Applications. Sobott, F., Ashcroft, A. E. (Eds). Royal Society of Chemistry.
- Chai, M. & Bleiholder, C. (2021). Structure-elucidation of human CCL5 by integrating trapped ion mobility spectrometry-mass spectrometry (TIMS-MS) with Structure Relaxation Approximation (SRA) analysis. International Journal of Mass Spectrometry, 469,116682.
- Bleiholder, C., Liu, F. C. (2019) A structure relaxation approximation (SRA) method to elucidate protein structures from ion mobility measurements. The Journal of Physical Chemistry B 123, 2756−2769.
- Young, M. N., & Bleiholder, C. (2017). Molecular Structures and Momentum Transfer Cross Sections: The Influence of the Analyte Charge Distribution. Journal of the American Society for Mass Spectrometry, 28, 619-627.
- Bleiholder, C. (2015) A local collision probability approximation for predicting momentum transfer cross sections. Analyst 14, 6804-6813.
- Bleiholder, C., Wyttenbach, T., Bowers, M. T. (2011) A novel projection approximation algorithm for the fast and accurate computation of molecular collision cross sections (I). Method. International Journal of Mass Spectrometry 308, 1-10.
Applications: Structural study of glycoproteins, carbohydrates, and chemokines by tandem-TIMS/MS
Heterogenous biological systems such as glycoproteins, carbohydrates and chemokines are often linked to diseases. Structural heterogeneity of these samples gives rise to many analytical challenges for more conventional biophysical tools.
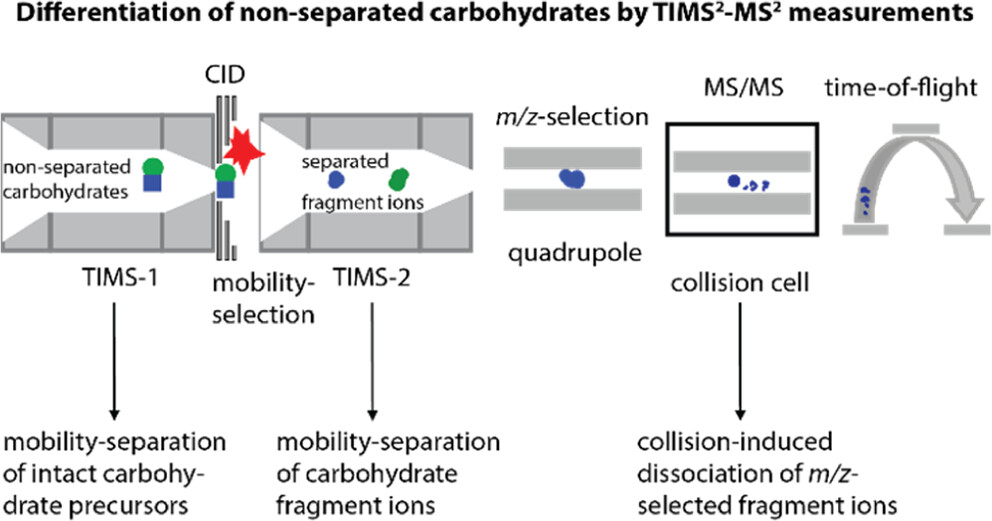
We exploit the ability of tandem-TIMS/MS to perform various operational workflows to study the quaternary, tertiary, and primary structures of glycoproteins. As shown for the glycoprotein avidin, tandem-TIMS/MS is able to retain the native-like tetrameric protein structure. Using collisional-induced dissociation experiments conducted between the two TIMS analyzers, we are able to identify subcomplexes of avidin. Further, combining two ion activation methods in tandem-TIMS/MS enables the top-down sequencing of avidin tetramers with a good sequence coverage. We further utilize TIMS2-MS2 workflows to differentiate and characterize isomeric, non-separable carbohydrates. In our ongoing research, we employ our second generation tandem-TIMS/MS equipped with a 213 nm laser for UV photodissociation, to examine more complex glycoprotein complexes and carbohydrates with highly diverse branching patterns.
Selected publications:
- Lee, J.; Chai, M.; & Bleiholder, C. (2023). Differentiation of isomeric, non-separable carbohydrates using tandem-trapped ion mobility spectrometry – mass spectrometry (tTIMS/MS). Analytical Chemistry 95 (2), 747–757.
- Liu, F. C., Kirk, S. R., Caldwell, K. A., Pedrete, T., Meier, F., & Bleiholder, C. (2022). Tandem Trapped Ion Mobility Spectrometry/Mass Spectrometry (tTIMS/MS) Reveals Sequence-Specific Determinants of Top-Down Protein Fragment Ion Cross Sections. Analytical Chemistry 94 (23), 8146–8155.
- Liu, F. C., Ridgeway, M E., Park, M. A., & Bleiholder, C. (2022). Tandem-trapped ion mobility spectrometry/mass spectrometry (tTIMS/MS): a promising analytical method for investigating heterogenous samples. Analyst 147, 2317-2337.
- Cropley, T. C., Chai, M., Liu, F. C., & Bleiholder, C. (2023). Perspective on the potential of tandem-ion mobility /mass spectrometry methods for structural proteomics applications. Frontiers in Analytical Science 3:1106752.
